A research team led by scientists with the Salk Institute for Biological Studies in California have applied the latest gene sequencing and mapping technology to take a closer look at the unintended consequences of the method most commonly used to create genetically engineered plants, including today’s herbicide-resistant varieties of corn, soybeans, sugarbeets, and cotton, as well as Bt-transgenic, insect-protected corn and cotton.
In a paper published this month in PLoS Genetics, the scientists closely examined how the tumor-inducing Agrobacterium tumefaciens bacteria can cause sometimes major, unintended changes to the genetic code of the target plant.
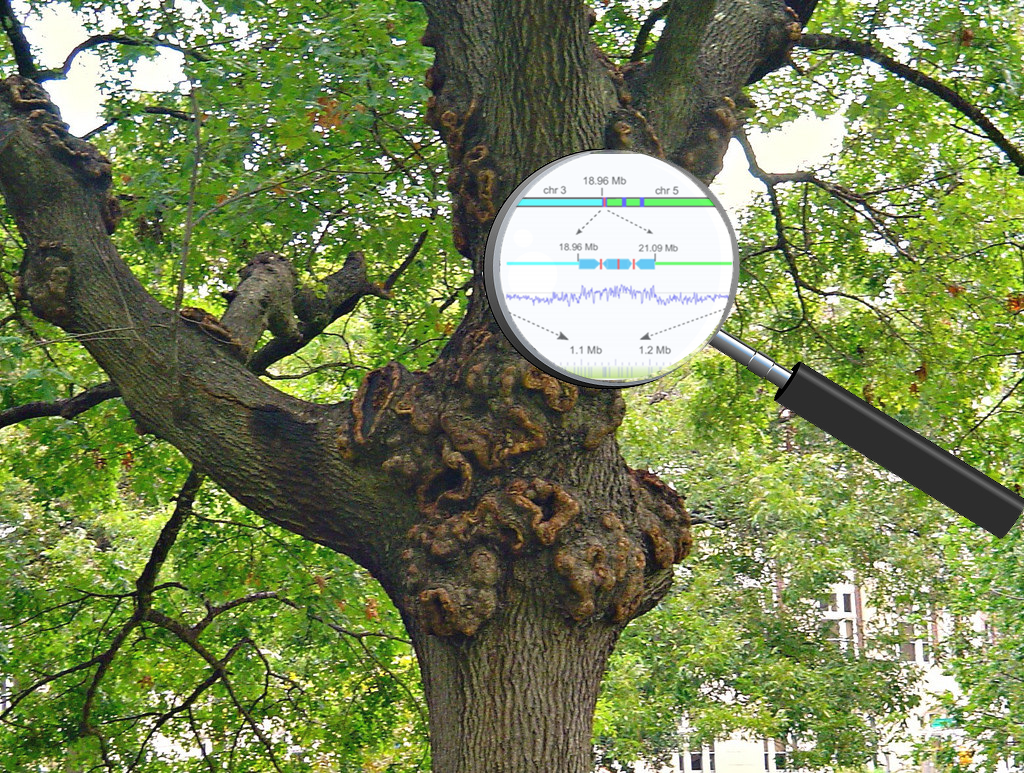
A. tumefaciens is considered the “workhorse” of the plant genetic-engineering field. The Agrobacterium genus includes bacteria that cause large bulges on branches and trunks of trees known as gall tumors (see image at top right). When GE-crop technology pioneers discovered decades ago that Agrobacterium bacteria transferred some of its DNA to host plants, it quickly became the go-to tool for the genetic engineering of plants.
Today, scientists use this unique capability of A. tumefaciens to transfer DNA into genetically-engineered plants, by attaching foreign DNA conferring a desirable trait to the natural DNA (called transfer-DNA or T-DNA) that moves from A. tumefaciens into the target, host plant that scientists are trying to transform.
Unfortunately though, this method for moving foreign DNA into plants results in “side-effects.” Such genetic-transformation miscues include:
- Expressing the foreign DNA in more than one place within a crop genome, or in the wrong place;
- Altering genetically-driven, signaling pathways that must work just right (and only when needed) to express certain proteins essential to normal plant growth and development, or to help the plant deal with stress; and/or
- Impacting the expression (when genes get turned on and off) of non-target genes that adversely impacts the fitness and health of the plant, or perhaps its composition, and hence nutritional quality.
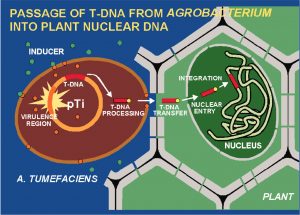
Remember – in an ideal world in which the genetic engineering of plants is “precise,” the T-DNA process would result in the insertion of just one, functional copy of the new genetic material, right where it belongs, and leave untouched all other genes within the plant’s genetic code.
Joseph Ecker is the head of the Genomic Analysis Laboratory at the Salk Institute and a co-author of the paper. He explains their objectives in a piece on the Salk Institute’s website:
“Biotech companies spend a lot of time and effort to characterize transgenic plants and disregard candidates with unwanted changes without understanding—from a basic biological perspective—why these changes may have occurred. Our new approach offers a way to better understand these effects and may help to speed up the process” (Salk News, 2019).
Until recently, such “unwanted changes” were simply disregarded and forgotten, like footage cut out of a film. However, Ecker and his team were able to use the latest, high-resolution genomic tools to take a closer look at how genetic transformation with A. tumefaciens impacts the target plant’s genetic code.
Their research “revealed that the plants had between one and seven distinct insertions or rearrangements in their genomes, ranging in size by almost tenfold…Gene insertions themselves showed a variety of patterns, with the inserted DNA fragment sometimes scrambled, inverted or even silenced.”
In other words, A. tumefaciens-driven genetic engineering is a far cry from precise, and is in fact inherently random and prone to unintended, complex genetic anomalies. And these unintended genetic changes matter – the team documented transformation results that “potentially changed the regulation or activation of other nearby genes” (Salk News, 2019).
The unintended consequences of genetic engineering are a recurrent focus of a number of “Hot Science” items on Hygeia. For a couple key examples, see this piece that presents the first clear evidence of “subtle metabolic changes” in GMO corn, or this story on a meta-analysis of genetic data on five different crops that documents wide variation between non-GMO and GMO maize, pumpkin, and rice.
This new Salk Institute study provides a “happy face” on the evolving capability of scientists to track the 800-pound gorilla in the closets of plant biotech laboratories across the country.
Hopefully this latest step forward in understanding the imprecision of first-generation biotech tools will give rise to greater care and forethought in shaping how gen-two tools like CRSPR and gene-editing are deployed. Understanding what is causing problems is, after all, always a useful step in solving them.
Sources:
Jupe, Florian; Rivkin, Angeline C; Michael, Todd P; Zander, Mark; Motley, S Timothy; Sandoval, Justin P; Slotkin, R Keith; Chen, Huaming; Castanon, Rosa; Nery, Joseph R; Ecker, Joseph R, “The complex architecture and epigenomic impact of plant T-DNA insertions,” PLoS Genetics, 2019, 15:e1007819, doi:10.1371/journal.pgen.1007819.
Salk News, “New Technologies Enable Better-Than-Ever Details on Genetically Modified Plants,” Epub Date: January 18, 2019, Access Date: January 28, 2019.

